About Us
Genetics plays an incredibly complex role in how neurodegenerative diseases like Alzheimer’s and Parkinson’s diseases appear and develop. Even small changes in the DNA sequence and structure can directly alter the protein product of a gene, or change how, when, and/or where a gene or a group of genes are expressed; these effects determine whether a disease will occur, when it happens, and the extent of its symptoms. Our research attempts to better understand the genetic processes underpinning age-related neurodegenerative diseases, in particular Alzheimer’s disease, related dementia, and Lewy body spectrum disorders.
Research
Our lab’s research focuses on the genetic factors and molecular mechanisms underlying age-related neurodegenerative diseases, in particular Alzheimer’s disease, related dementia, and Lewy body spectrum disorders. Our goal is to identify the precise causal and functional genetic variants underlying GWAS discoveries, and to understand their molecular mechanisms of action and the biological pathways through which they exert their pathogenic effects. To this end, we develop innovative in silico, in vitro, and in vivo approaches to study noncoding variants, specifically short structural variants, their corresponding trans factors, and their impact on the regulation of gene expression and splicing in the context of neurodegenerative phenotypes. Our research program has translational applications, informing clinical studies and supporting the development of novel genetic biomarkers, therapeutic targets, and ‘next generation’ gene therapy approaches.
Developing isogenic hiPSC-derived model systems for studying age-related neurodegenerative diseases
Modeling human age-related neurodegenerative diseases (NDDs), including synucleinopathies, by using human induced pluripotent stem cells (hiPSCs) leverages the use of cell culture systems in research of neurodegenerative diseases. The hiPSC system has several advantages crucial to accomplishing our research goals. First, as an isogenic system, it allows us to examine the direct effects of specific genetic variants on a common human genetic background, in a cell-specific manner. Second, it enables us to perform comprehensive assessments of molecular phenotypes and cellular functions using living cells.
We have developed isogenic hiPSC-derived aged cholinergic and dopaminergic neurons, and astrocytes involved in age-related NDDs, including AD, PD and DLB. These 2D model systems are suitable for investigating cell-specific disease mechanisms and genetic etiologies, and to validate causal genetic factors.

A deeper understanding of the interplay between brain cell types is feasible with novel 3D co-culture systems. The isogenic hiPSC-derived 3D triculture technique recapitulates the brain microenvironment in vitro and enables to study the interactions between the different brain cell types. Working with the Bursac group, we initiate the development of 3D triculture systems composed of various combinations of isogenic neurons, astrocytes, and microglia as human-based disease models for exploring the mechanistic complexity of NDDs.
Additionally, in collaboration with the Kantor and Akkari research teams, our overarching goal is to translate the mechanistic understanding to clinical applications. The hiPSC-derived models facilitate the validation of therapeutic targets and the establishment of proof-of-concept for next generation gene-therapy approaches, including CRISPR/Cas (genome and epigenome editing) and antisense oligonucleotide (ASO) technologies, as a precision medicine for individuals at high risk for NDDs due to gene dysregulation.

Decoding the genetic architecture of late-onset Alzheimer’s disease using single-cell multi-‘omics
Identifying neuron-, astrocyte- and microglia-specific alterations in regulatory elements, the epigenome landscape, and gene expression profiles between normal and LOAD-pathological stages will advance the identification of target genes within the LOAD-associated loci.
In collaboration with the Crawford and the Ashley-Koch labs we investigate chromatin accessibility and gene expression profiles in affected brain tissues from LOAD patients. The heterogeneity of brain tissue makes it difficult to assess molecular characteristics of individual cell types. Frozen tissue presents additional challenges, as intact cell bodies are more difficult to isolate than intact nuclei. Therefore, we used fluorescence-activated nuclei sorting (FANS) to distinguish between neuronal and non-neuronal nuclei. Unexpectedly, we detected LOAD-specific differential chromatin accessibility in non-neuronal cells from female but not male subjects. This has prompted further investigation to determine the specific non-neuronal cell types underlying this sex-specific genetic regulatory landscape.
To interrogate cellular diversity in the brain at an even higher resolution, our lab employs the 10X Genomics microfluidics system to generate single-nuclei (sn) barcoded libraries for RNA-sequencing (seq) and ATAC-seq. We perform snRNA-seq and snATAC-seq in parallel from the same sample, such that chromatin accessibility and gene expression profiles can be assessed from the same pool of nuclei. We utilize advanced data science methods to process and analyze these large datasets and generate cell type-specific clusters. Ultimately, we are able to characterize and integrate cell subtype-specific multi-omic profiles to identify regulatory elements implicated in LOAD pathogenesis and progression.
Bioinformations pipeline integrating functional genomic datasets to decipher LOAD-GWAS discoveries to causal genes and regulatory variants
In the post genome-wide-association-studies (GWAS) era we are shifting gears toward translation of genetic disease loci to molecular mechanisms of pathogenesis. Large multi-center GWAS have found associations between >30 genomic loci and late-onset Alzheimer’s disease (LOAD), and candidate genes were inferred by the proximity to the associated-SNP. However, the precise target genes within the LOAD-associated genomic regions and the causal variants affecting LOAD-risk have yet to be uncovered. In collaboration with Dr. Lutz, we developed a bioinformatics pipeline based on functional genomic datasets to advance the interpretation of Alzheimer's disease GWAS discoveries. This pipeline can be applied to other complex genetic diseases.
Pleiotropy between LOAD and Neuropsychiatric conditions
Several key observations have provided evidence for the interrelationship between Alzheimer’s disease (AD) and neuropsychiatric symptoms (NPS). Neuropsychiatric conditions have been shown to increase the risk for developing dementia, and patients with AD frequently manifest comorbid NPS with wide heterogeneity. This evidence suggests shared genetic etiology and intersecting pathways between AD and neuropsychiatric conditions, which contributes to the comorbid phenotypes of NPS, such as depression and anxiety. In collaboration with Drs. Lutz, Williamson, and Sheng, we have been testing the hypothesis that genetic variability, epigenetic signatures, and gene expression changes in specific genes are common between AD and neuropsychiatric conditions, and that these shared molecular phenotypes are responsible, at least in part, for the heterogeneity of NPS in AD.
In particular, we have been studying two neuropsychiatric disorders, Major Depressive Disorder (MDD) and Post-traumatic Stress Disorder (PTSD). For both we found a moderate enrichment for SNPs associated with LOAD across increasingly stringent levels of significance with the MDD and the PTSD GWAS findings. Association analysis identified numerous pleiotropic risk-loci, with the primary and most significant SNPs positioned in two regions of chromosome 11 (SPI1 gene and MS4A genes cluster). Functional genomic studies, including eQTLs and pathway analyses, pointed to the immune response as a common pathway.
The broader contribution of SNCA gene to the wide spectrum of Lewy Body disorders
A neuropathological hallmark of a group of neurodegenerative diseases, known as human synucleinopathies, is the presence of intracellular protein aggregates named Lewy bodies (LBs) and Lewy neurites (LNs). The alpha-synuclein protein is a major component of LBs. Parkinson’s disease (PD) is the archetypal human synucleinopathy and has been studied extensively. Genome-wide association studies (GWAS) have implicated the alpha-synuclein gene (SNCA) in PD, DLB and MSA. Several studies in cell-culture and animal models reported that overexpression of wild-type SNCA can be toxic and may lead to cell death. Moreover, we documented that elevated SNCA expression in patients correlated with Lewy body presentation.
We utilize a multi-faceted strategy in our experiments to decipher the role of SNCA in health and disease, whether that be LB pathologies in general or in a particular disease within the spectrum.
One of the leading projects in our lab aims to uncover the molecular mechanisms regulating SNCA expression (such as transcription, RNA metabolism), and any genetic variability that impinges on this regulation. Another more recent study in our lab suggested a role for SNCA in the nucleus in the context of aging. Specifically, we found that when SNCA is mutated or overexpressed, it disrupts the nucleocytoplasmic transport leading to impairment of nuclear function and exacerbation of aging.
The role of TOMM40-APOE genomic region in late-onset Alzheimer's disease
APOE ε4 is the strongest and most replicated genetic risk factor for late-onset Alzheimer’s disease (LOAD). APOE is located on chromosome 19 (19q13.32) in the tight gene cluster TOMM40-APOE-APOC1-APOC4-APOC2 that exhibits strong linkage disequilibrium (LD). Genome wide association studies (GWAS) reported that the strongest association signal (by a wide margin) was also in the APOE LD region. This top association signal was attributed to the APOE ε4 haplotype; however, it has been acknowledged that other genetic factors within this LD block may also explain the strong genome-wide significant signal and contribute, in part, to disease risk attributed to this genomic region. This work has been carried out in collaborations with the late Dr. Allen Roses and team, and Drs. Lutz and Gottschalk.
We investigated the TOMM40-APOE genomic region associated with the risk and age of onset of LOAD to determine the functional consequences of a highly polymorphic, intronic polyT within this region (rs10524523) discovered by Roses et al. The TOMM40-polyT was associated with LOAD age of onset and other disease related endophenotypes, and we have identified an association with cognitive performance in normal aging. Specifically, we have been studying the regional regulatory effect of the TOMM40-polyT site on gene expression. In these investigations, we are using multiple model systems, including human brain tissues, cell-based reporter systems, mESCs, and humanized mouse models. We have found that the polyT acts as a regional transcriptional regulator of TOMM40 and APOE genes, suggesting that the TOMM40-polyT locus may contribute to LOAD susceptibility by modulating TOMM40 and/or APOE mRNA expression levels. We have also manipulated the transcriptional effects on TOMM40 and APOE-C1 genes in vitro using two approaches: knockdown of PPARgamma with shRNA, and treatment with pharmacological agents that are known PPARgamma agonists, particularly low doses of the Pioglitazone and Rosiglitazone compounds.
'Next Generation' gene therapy approaches for age-related neurodegenerative diseases caused by dysregulation of key genes
Dysregulation of gene expression has been implicated in the etiology of several age-related neurodegenerative diseases. Translating the knowledge of the mechanisms involved in the regulation of firmly established key-genes in disease, such as SNCA in PD and APOE in LOAD, will facilitate next generation drug discovery based on targeted manipulation of gene expression in a fine-tuned manner. This therapeutic strategy is efficient for fine modulation of gene expression to restore physiological levels of critical proteins. In collaboration with the Kantor lab, we developed a novel strategy to intervene with transcriptional regulation programs of SNCA and APOE, based on targeted epigenome editing. Specifically, we have established a system for targeted DNA-methylation editing that comprises an all-in-one lentiviral vector, for the delivery of CRISPR/dCas9 fused with the catalytic domain of DNA-methyltransferase 3A (DNMT3A).
Applying this novel CRISPR/dCas9-based technology offers an unprecedented tool to modify a particular epigenetic mark. Specifically, this technology resulted in effective fine-tuned reduction of SNCA expression levels in hiPSC-derived neurons from a PD patient that was sufficient for reversing PD-associated phenotypic perturbations. These proof-of-concept experiments provide the foundation and validation for advancing this novel epigenetic editing-based system towards a PD therapeutic strategy applicable in a clinical setting. Our innovative platform that allows for fine-tuned regulation of gene expression would be highly appealing and translational for developing ‘next generation drugs’ as prevention and/or disease modifying interventions for PD, Alzheimer’s disease and other neurodegenerative diseases and pathologies associated with dysregulation of gene expression.
Our Team
Graduates and former employees of the Chiba-Falek laboratory have gone on to excel both in clinical medicine and laboratory research. The list below notes former members of our laboratory.
Pierce Augusti
Undergraduate Student
Sneha Chandrashekar
Summer Student | Genome Science and Medicine
Zhaohui Man
Bioinformation I
Christian Frost
Research Technician II
Justin Fraser
Summer Student | Genome Science and Medicine
Wiljeen Paul
Rotation Student
Webb Pierson | Staff Spotlight
Volunteer
Peter Nicholls, MD
Research Scientist
Julio Barrera
Laboratory Manager
Julia Gamache, PhD
Postdoctoral Associate
Jeison Valencia
Research Technician II
Suraj Upadhya
Undergraduate Student
Cordelia Hume
Undergraduate Student
Angela Wei
Undergraduate Student
Anna Yang
Undergraduate Student
Chesney Birshing
Summer Student
Delaney Hill
Summer Student
Danielle Chipman
Undergraduate Student
Kennedy Lofton
Undergraduate Student
Gabriella MacDougall, PhD
Postdoctoral Associate
Daniel Sprague
Undergraduate Student
Jeffrey Gu
Undergraduate Student
Vivian Chen
Undergraduate Student
Manish Kumar
Undergraduate Student
Dominic Tringali
Research Technician
Young Yun
Research Technician
Lidia Tagliafierro, PhD
Postdoctoral Associate
Ivana Premasinghe
Undergraduate Student
Ahlia Sriskanda
Research technician
George Crawley, IV
Undergraduate Student
Madison Zamora
Undergraduate Student
Kathy Ziyue Dai
Undergraduate Student
Omolara-Chinue Glenn
Research technician
Shobana Subramanian
Undergraduate Student
Kirsten Bonawitz
Undergraduate Student
Haylee Bergstrom
Undergraduate Student
Taylor Novice
Undergraduate Student
Christina He
Undergraduate Student
Natalie Miller
Undergraduate Student
Jawara Allen
Undergraduate Student
Sunita Saith
Undergraduate Student
Jennifer Byrne
Research Technician
Laura Saucier
Undergraduate Student
Recent Publications
- Neuronal-type-specific epigenome editing to decrease SNCA expression: Implications for precision medicine in synucleinopathies. Sun Z, Kantor B, Chiba-Falek O. Mol Ther Nucleic Acids. 2023 Nov 24;35(1):102084. doi: 10.1016/j.omtn.2023.102084.
- Integrative single-nucleus multi-omics analysis prioritizes candidate cis and trans regulatory networks and their target genes in Alzheimer's disease brains. Gamache J, Gingerich D, Shwab EK, Barrera J, Garrett ME, Hume C, Crawford GE, Ashley-Koch AE, Chiba-Falek O. Cell Biosci. 2023 Oct 3;13(1):185. doi: 10.1186/s13578-023-01120-5.
- The APOE-TOMM40 Humanized Mouse Model: Characterization of Age, Sex, and PolyT Variant Effects on Gene Expression. Gottschalk WK, Mahon S, Hodgson D, Barrera J, Hill D, Wei A, Kumar M, Dai K, Anderson L, Mihovilovic M, Lutz MW, Chiba-Falek O. J Alzheimers Dis. 2023;94(4):1563-1576. doi: 10.3233/JAD-230451.
- Bioinformatics pipeline to guide post-GWAS studies in Alzheimer's: A new catalogue of disease candidate short structural variants. Lutz MW, Chiba-Falek O. Alzheimers Dement. 2023 Sep;19(9):4094-4109. doi: 10.1002/alz.13168.
- Differential Gene Expression and DNA Methylation in the Risk of Depression in LOAD Patients. Upadhya S, Gingerich D, Lutz MW, Chiba-Falek O. Biomolecules. 2022 Nov 12;12(11):1679. doi: 10.3390/biom12111679.
- Polygenic Risk Score Effectively Predicts Depression Onset in Alzheimer's Disease Based on Major Depressive Disorder Risk Variants. Upadhya S, Liu H, Luo S, Lutz MW, Chiba-Falek O. Front Neurosci. 2022 Mar 8;16:827447. doi: 10.3389/fnins.2022.827447.
- Bioinformatics pipeline to guide late-onset Alzheimer's disease (LOAD) post-GWAS studies: Prioritizing transcription regulatory variants within LOAD-associated regions. Lutz MW, Chiba-Falek O. Alzheimers Dement (N Y). 2022 Feb 23;8(1):e12244. doi: 10.1002/trc2.12244.
- Sex dependent glial-specific changes in the chromatin accessibility landscape in late-onset Alzheimer's disease brains. Barrera J, Song L, Gamache JE, Garrett ME, Safi A, Yun Y, Premasinghe I, Sprague D, Chipman D, Li J, Fradin H, Soldano K, Gordân R, Ashley-Koch AE, Crawford GE, Chiba-Falek O. Mol Neurodegener. 2021 Aug 24;16(1):58. doi: 10.1186/s13024-021-00481-0.
- Identifying nootropic drug targets via large-scale cognitive GWAS and transcriptomics. Lam M, Chen CY, Ge T, Xia Y, Hill DW, Trampush JW, Yu J, Knowles E, Davies G, Stahl EA, Huckins L, Liewald DC, Djurovic S, Melle I, Christoforou A, Reinvang I, DeRosse P, Lundervold AJ, Steen VM, Espeseth T, Räikkönen K, Widen E, Palotie A, Eriksson JG, Giegling I, Konte B, Hartmann AM, Roussos P, Giakoumaki S, Burdick KE, Payton A, Ollier W, Chiba-Falek O, Koltai DC, Need AC, Cirulli ET, Voineskos AN, Stefanis NC, Avramopoulos D, Hatzimanolis A, Smyrnis N, Bilder RM, Freimer NB, Cannon TD, London E, Poldrack RA, Sabb FW, Congdon E, Conley ED, Scult MA, Dickinson D, Straub RE, Donohoe G, Morris D, Corvin A, Gill M, Hariri AR, Weinberger DR, Pendleton N, Bitsios P, Rujescu D, Lahti J, Le Hellard S, Keller MC, Andreassen OA, Deary IJ, Glahn DC, Huang H, Liu C, Malhotra AK, Lencz T. Neuropsychopharmacology. 2021 Sep;46(10):1788-1801. doi: 10.1038/s41386-021-01023-4.
- Cell-Type Specific Changes in DNA Methylation of SNCA Intron 1 in Synucleinopathy Brains. Gu J, Barrera J, Yun Y, Murphy SK, Beach TG, Woltjer RL, Serrano GE, Kantor B, Chiba-Falek O. Front Neurosci. 2021 Apr 28;15:652226. doi: 10.3389/fnins.2021.652226.
- APOE: The New Frontier in the Development of a Therapeutic Target towards Precision Medicine in Late-Onset Alzheimer's. Yang A, Kantor B, Chiba-Falek O. Int J Mol Sci. 2021 Jan 27;22(3):1244. doi: 10.3390/ijms22031244.
- The Path to Progress Preclinical Studies of Age-Related Neurodegenerative Diseases: A Perspective on Rodent and hiPSC-Derived Models. MacDougall G, Brown LY, Kantor B, Chiba-Falek O. Mol Ther. 2021 Mar 3;29(3):949-972. doi: 10.1016/j.ymthe.2021.01.001.
- Good problems to have? Policy and societal implications of a disease-modifying therapy for presymptomatic late-onset Alzheimer's disease. Angrist M, Yang A, Kantor B, Chiba-Falek O. Life Sci Soc Policy. 2020 Oct 12;16(1):11. doi: 10.1186/s40504-020-00106-2.
- The mechanistic role of alpha-synuclein in the nucleus: impaired nuclear function caused by familial Parkinson's disease SNCA mutations. Chen V, Moncalvo M, Tringali D, Tagliafierro L, Shriskanda A, Ilich E, Dong W, Kantor B, Chiba-Falek O. Hum Mol Genet. 2020 Nov 4;29(18):3107-3121. doi: 10.1093/hmg/ddaa183.
- Gene-Editing Technologies Paired With Viral Vectors for Translational Research Into Neurodegenerative Diseases. Rittiner JE, Moncalvo M, Chiba-Falek O, Kantor B. Front Mol Neurosci. 2020 Aug 12;13:148. doi: 10.3389/fnmol.2020.00148.
- Sex-dependent effect of APOE on Alzheimer's disease and other age-related neurodegenerative disorders. Gamache J, Yun Y, Chiba-Falek O. Dis Model Mech. 2020 Aug 27;13(8):dmm045211. doi: 10.1242/dmm.045211.
- Shared genetic etiology underlying late-onset Alzheimer's disease and posttraumatic stress syndrome. Lutz MW, Luo S, Williamson DE, Chiba-Falek O. Alzheimers Dement. 2020 Sep;16(9):1280-1292. doi: 10.1002/alz.12128.
- Shared Genetic Etiology Underlying Alzheimer's Disease and Major Depressive Disorder. Lutz MW, Sprague D, Barrera J, Chiba-Falek O. Transl Psychiatry. 2020 Mar 9;10(1):88. doi: 10.1038/s41398-020-0769-y.
- Pleiotropic Meta-Analysis of Cognition, Education, and Schizophrenia Differentiates Roles of Early Neurodevelopmental and Adult Synaptic Pathways. Lam M, Hill WD, Trampush JW, Yu J, Knowles E, Davies G, Stahl E, Huckins L, Liewald DC, Djurovic S, Melle I, Sundet K, Christoforou A, Reinvang I, DeRosse P, Lundervold AJ, Steen VM, Espeseth T, Räikkönen K, Widen E, Palotie A, Eriksson JG, Giegling I, Konte B, Hartmann AM, Roussos P, Giakoumaki S, Burdick KE, Payton A, Ollier W, Chiba-Falek O, Attix DK, Need AC, Cirulli ET, Voineskos AN, Stefanis NC, Avramopoulos D, Hatzimanolis A, Arking DE, Smyrnis N, Bilder RM, Freimer NA, Cannon TD, London E, Poldrack RA, Sabb FW, Congdon E, Conley ED, Scult MA, Dickinson D, Straub RE, Donohoe G, Morris D, Corvin A, Gill M, Hariri AR, Weinberger DR, Pendleton N, Bitsios P, Rujescu D, Lahti J, Le Hellard S, Keller MC, Andreassen OA, Deary IJ, Glahn DC, Malhotra AK, Lencz T. Am J Hum Genet. 2019 Aug 1;105(2):334-350. doi: 10.1016/j.ajhg.2019.06.012.
- The Landscape of SNCA Transcripts Across Synucleinopathies: New Insights From Long Reads Sequencing Analysis. Tseng E, Rowell WJ, Glenn OC, Hon T, Barrera J, Kujawa S, Chiba-Falek O. Front Genet. 2019 Jul 9;10:584. doi: 10.3389/fgene.2019.00584. eCollection 2019.
- Bioinformatics strategy to advance the interpretation of Alzheimer's disease GWAS discoveries: The roads from association to causation. Lutz MW, Sprague D, Chiba-Falek O. Alzheimers Dement. 2019 Jun 28. pii: S1552-5260(19)30121-9. doi: 10.1016/j.jalz.2019.04.014.
- Lentiviral Vector Platform for the Efficient Delivery of Epigenome-editing Tools into Human Induced Pluripotent Stem Cell-derived Disease Models. Tagliafierro L, Ilich E, Moncalvo M, Gu J, Sriskanda A, Grenier C, Murphy SK, Chiba-Falek O, Kantor B. J Vis Exp. 2019 Mar 29;(145). doi: 10.3791/59241.
- Multiplication of the SNCA locus exacerbates neuronal nuclear aging. Tagliafierro L, Zamora ME, Chiba-Falek O. Hum Mol Genet. 2019 Feb 1;28(3):407-421. doi: 10.1093/hmg/ddy355.
- Toward deciphering the mechanistic role of variations in the Rep1 repeat site in the transcription regulation of SNCA gene. Afek A, Tagliafierro L, Glenn OC, Lukatsky DB, Gordan R, Chiba-Falek O. Neurogenetics. 2018 Aug;19(3):135-144. doi: 10.1007/s10048-018-0546-8. Epub 2018 May 5.
- Probing the role of PPARγ in the regulation of late-onset Alzheimer's disease-associated genes. Barrera J, Subramanian S, Chiba-Falek O. PLoS One. 2018 May 3;13(5):e0196943. doi: 10.1371/journal.pone.0196943. eCollection 2018.
- The effects of the TOMM40 poly-T alleles on Alzheimer's disease phenotypes. Chiba-Falek O, Gottschalk WK, Lutz MW. Alzheimers Dement. 2018 May;14(5):692-698. doi: 10.1016/j.jalz.2018.01.015. Epub 2018 Mar 7.
- Genetic analysis of α-synuclein 3' untranslated region and its corresponding microRNAs in relation to Parkinson's compared to dementia with Lewy bodies. Tagliafierro L, Glenn OC, Zamora ME, Beach TG, Woltjer RL, Lutz MW, Chiba-Falek O. Alzheimers Dement. 2017 Apr 18. pii: S1552-5260(17)30136-X. doi: 10.1016/j.jalz.2017.03.001.
- Structural variants in SNCA gene and the implication to synucleinopathies. Chiba-Falek O. Curr Opin Genet Dev. 2017 Jun;44:110-116. doi: 10.1016/j.gde.2017.01.014. Epub 2017 Mar 2. Review.
- The effects of PPARγ on the regulation of the TOMM40-APOE-C1 genes cluster. Subramanian S, Gottschalk WK, Kim SY, Roses AD, Chiba-Falek O. Biochim Biophys Acta. 2017 Mar;1863(3):810-816. doi: 10.1016/j.bbadis.2017.01.004.
- GWAS meta-analysis reveals novel loci and genetic correlates for general cognitive function: a report from the COGENT consortium. Trampush JW, Yang ML, Yu J, Knowles E, Davies G, Liewald DC, Starr JM, Djurovic S, Melle I, Sundet K, Christoforou A, Reinvang I, DeRosse P, Lundervold AJ, Steen VM, Espeseth T, Räikkönen K, Widen E, Palotie A, Eriksson JG, Giegling I, Konte B, Roussos P, Giakoumaki S, Burdick KE, Payton A, Ollier W, Horan M, Chiba-Falek O, Attix DK, Need AC, Cirulli ET, Voineskos AN, Stefanis NC, Avramopoulos D, Hatzimanolis A, Arking DE, Smyrnis N, Bilder RM, Freimer NA, Cannon TD, London E, Poldrack RA, Sabb FW, Congdon E, Conley ED, Scult MA, Dickinson D, Straub RE, Donohoe G, Morris D, Corvin A, Gill M, Hariri AR, Weinberger DR, Pendleton N, Bitsios P, Rujescu D, Lahti J, Le Hellard S, Keller MC, Andreassen OA, Deary IJ, Glahn DC, Malhotra AK, Lencz T. Mol Psychiatry. 2017 Mar;22(3):336-345. doi: 10.1038/mp.2016.244. Epub 2017 Jan 17.
- Gene Expression Analysis of Neurons and Astrocytes Isolated by Laser Capture Microdissection from Frozen Human Brain Tissues. Tagliafierro L, Bonawitz K, Glenn OC, Chiba-Falek O. Front Mol Neurosci. 2016 Aug 18;9:72. doi: 10.3389/fnmol.2016.00072.
- The SSV Evaluation System: A Tool to Prioritize Short Structural Variants for Studies of Possible Regulatory and Causal Variants. Saul R, Lutz MW, Burns DK, Roses AD, Chiba-Falek O. Hum Mutat. 2016 Sep;37(9):877-83. doi: 10.1002/humu.23023.
- The Role of Upregulated APOE in Alzheimer's Disease Etiology. Gottschalk WK, Mihovilovic M, Roses AD, Chiba-Falek O. J Alzheimers Dis Parkinsonism. 2016 Mar;6(1). pii: 209.
- Structural variants can be more informative for disease diagnostics, prognostics and translation than current SNP mapping and exon sequencing. Roses AD, Akkari PA, Chiba-Falek O, Lutz MW, Gottschalk WK, Saunders AM, Saul B, Sundseth S, Burns D. Expert Opin Drug Metab Toxicol. 2016;12(2):135-47. doi: 10.1517/17425255.2016.1133586.
- A cytosine-thymine (CT)-rich haplotype in intron 4 of SNCA confers risk for Lewy body pathology in Alzheimer's disease and affects SNCA expression. Lutz MW, Saul R, Linnertz C, Glenn OC, Roses AD, Chiba-Falek O. Alzheimers Dement. 2015 Oct;11(10):1133-43. doi: 10.1016/j.jalz.2015.05.011.
- The Broad Impact of TOM40 on Neurodegenerative Diseases in Aging. Gottschalk WK, Lutz MW, He YT, Saunders AM, Burns DK, Roses AD, Chiba-Falek O. J Parkinsons Dis Alzheimers Dis. 2014 Nov;1(1). pii: 12.
Collaborators
Our laboratory welcomes collaborating researchers, from within Duke as well as universities around the world. The following experts are some of the people we work with to advance our knowledge of the genetic factors and molecular mechanisms underlying age-related neurodegenerative diseases.
- William Kirby Gottschalk, PhD - Duke University School of Medicine, Department of Neurology
- Michael Lutz, PhD – Duke University School of Medicine, Department of Neurology
- Allison Ashley-Koch, PhD - Duke University School of Medicine, Duke Molecular Physiology Institute
- Yarui Diao, PhD - Duke University School of Medicine, Department of Cell Biology
- Raluca Gordân, PhD - Duke University School of Medicine, Biostatistics & Bioinformatics and Computer Science and Duke Center for Genomic and Computational Biology
- Greg Crawford, PhD - Duke University School of Medicine, Department of Pediatrics and Duke Center for Genomic and Computational Biology
- Susan K. Murphy, PhD - Duke University School of Medicine, Department of Obstetrics and Gynecology
- Boris Kantor, PhD - Duke University School of Medicine, Department of Neurobiology
- Thomas Beach, MD, PhD - Banner Sun Health Research Institute - Sun City, Arizona
- Anthony Akkari, PhD, Head, Motor Neuron Disease Genetics and Therapeutics Research, Perron Institute, Western Australia
- Randy Woltjer, MD, PhD - Oregon Health and Science University - Portland, Oregon
- David Lukatsky, PhD – Ben Gurion University, Israel
- Robert Saul, PhD - Polymorphic DNA Technologies, Inc, California
- Steve Wilton, PhD - Perron Institute Director, Foundation Chair Molecular Therapy
- Ziv Gan-Or, MD, PhD - McGill University, Department of Neurology - Montréal, QC
Photos
Justin Fraser, summer student from the GCB Summer Scholars Program in Genome Sciences & Medicine, presents his poster in July 2024.

Members of our translational research teams enjoy a holiday potluck dinner in this photo. Check out more photos of this holiday event here.
Chensey Birshing shares her research poster, "In vivo validitation of an α-synuclein-targeted epigenome therapy for Parkinson's disease" in summer of 2023.

The Chiba Falek lab celebrates during a 2023 team visit to Urban Axes.

Lab members Delila Hodgson, Christan Frost, Daniel Gingerich, and Webb Pierson (left from right) show off their axe throwing skills during a team visit to Urban Axes.

Graduating Duke University seniors Suraj Upadhya (left) and Scott Mahon (right) share their research during recent poster sessions.

The Chiba Falek team enjoys a holiday luncheon at JuJu restaurant, December 2022
Students Anna Yang and Angela Wei celebrate during the Chiba Falek team's 2022 end-of-year celebration dinner at Juju in Durham.

Members of the Chiba-Falek lab enjoy time together during the 2020 Translational Brain Sciences Poster session.
Daniel Sprague presents his research at Duke Science and Society's Huang Fellows Program Poster Session

George Crawley IV presents his research at the Duke Center for Applied Genomics & Precision Medicine's Summer Scholars Poster Presentation.

Members of the Chiba Falek lab pose during the Senior Undergraduate Neuroscience poster session for Graduation with Distinction in April 2017.
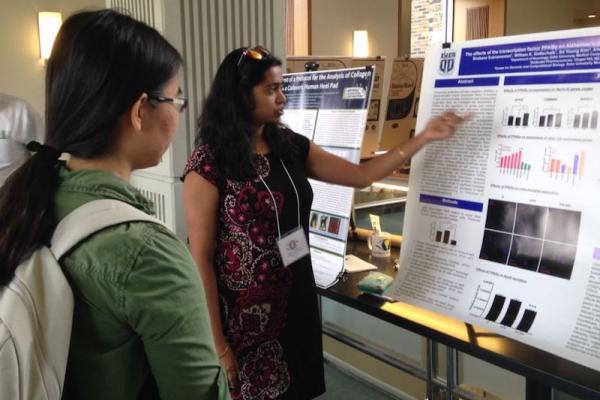

Shobana Subramanian explains her research project during the 2017 Duke Visible Thinking Poster Session.

Members of the Chiba-Falek team enjoy a night out to celebrate a fully year of work.
Kirsten Bonawitz explains her research project during the 2017 Duke Visible Thinking Poster Session.

Members of the Chiba-Falek lab pose outside of their home in the Bryan Research Building.

The Chiba Falek lab celebrates a Alzheimer’s Association International Research Grant for Lidia Tagliafierro, PhD (red, second from left).








